 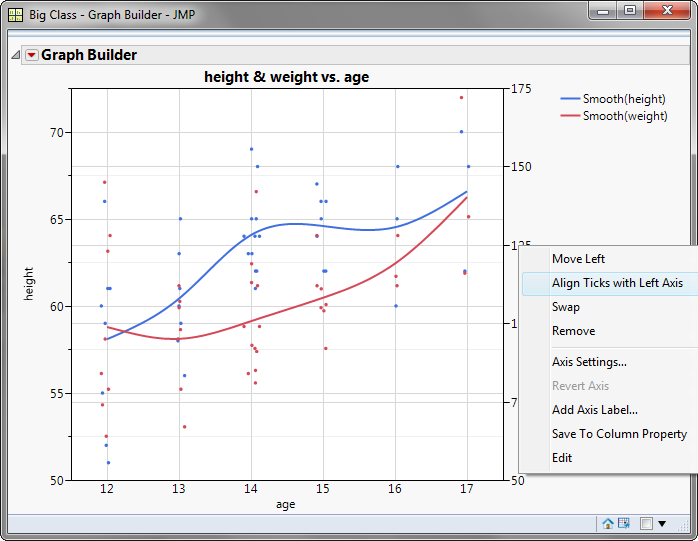
lmMod_bc <- lm(dist_new ~ speed, data=cars) Lets build the model and check for heteroscedasticity. The transformed data for our new regression model is ready. cars <- cbind(cars, dist_new=predict(distBCMod, cars$dist)) # append the transformed variable to cars Lets now apply it on car$dist and append it to a new dataframe. The model for creating the box-cox transformed variable is ready. distBCMod <- caret::BoxCoxTrans(cars$dist) Often, doing a box-cox transformation of the Y variable solves the issue, which is exactly what I am going to do now. Box-Cox transformationīox-cox transformation is a mathematical transformation of the variable to make it approximate to a normal distribution. Lets now hop on to Box-Cox transformation. Though is this not recommended, it is an approach you could try out if all available options fail. The solutions is, for starters, you could use the mean value of residuals for all observations in test data. With a model that includes residuals (as X) whose future actual values are unknown, you might ask what will be the value of the new predictor (i.e. However, one option I might consider trying out is to add the residuals of the original model as a predictor and rebuild the regression model. Since we have no other predictors apart from “speed”, I can’t show this method now. Variable transformation such as Box-Cox transformation. NCV Test car::ncvTest(lmMod) # Breusch-Pagan testĬhisquare = 4.650233 Df = 1 p = 0.03104933īoth these test have a p-value less that a significance level of 0.05, therefore we can reject the null hypothesis that the variance of the residuals is constant and infer that heteroscedasticity is indeed present, thereby confirming our graphical inference. For this purpose, there are a couple of tests that comes handy to establish the presence or absence of heteroscedasticity – The Breush-Pagan test and the NCV test.īreush Pagan Test lmtest::bptest(lmMod) # Breusch-Pagan test Sometimes you may want an algorithmic approach to check for heteroscedasticity so that you can quantify its presence automatically and make amends. So, the inference here is, heteroscedasticity exists. If there is absolutely no heteroscedastity, you should see a completely random, equal distribution of points throughout the range of X axis and a flat red line.īut in our case, as you can notice from the top-left plot, the red line is slightly curved and the residuals seem to increase as the fitted Y values increase. The top-left is the chart of residuals vs fitted values, while in the bottom-left one, it is standardised residuals on Y axis. The plots we are interested in are at the top-left and bottom-left. Graphical method par(mfrow=c(2,2)) # init 4 charts in 1 panel Now that the model is ready, there are two ways to test for heterosedasticity: lmMod <- lm(dist ~ speed, data=cars) # initial model Lets first build the model using the lm() function. I am going to illustrate this with an actual regression model based on the cars dataset, that comes built-in with R. This would result in an inefficient and unstable regression model that could yield bizarre predictions later on. The reason is, we want to check if the model thus built is unable to explain some pattern in the response variable \(Y\), that eventually shows up in the residuals. It is customary to check for heteroscedasticity of residuals once you build the linear regression model. Why is it important to check for heteroscedasticity? This process is sometimes referred to as residual analysis. In this post, I am going to explain why it is important to check for heteroscedasticity, how to detect it in your model? If is present, how to make amends to rectify the problem, with example R codes.  
In simpler terms, this means that the variance of residuals should not increase with fitted values of response variable. One of the important assumptions of linear regression is that, there should be no heteroscedasticity of residuals. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
